Special Offers
100% Performance Guaranteed
Key Specifications Table
| Species Reactivity | Key Applications |
|---|---|
| H, Vrt | WB, IP, EMSA, ChIP |
| Description | |
|---|---|
| Catalogue Number | 17-10136 |
| Trade Name |
|
| Description | ChIPAb+ Dimethyl-Histone H3 (Lys36) - ChIP Validated Antibody and Primer Set |
| Overview | All ChIPAb+ antibodies are individually validated for chromatin precipitation, every lot, every time. Each ChIPAb+ antibody set includes control primers (tested every lot by qPCR) to biologically validate your IP results in a locus-specific context. The qPCR protocol and primer sequences are provided, allowing researchers to validate ChIP protocols when using our antibody in their chromatin context. Each set also includes a negative control antibody to ensure specificity of the ChIP reaction. The ChIPAb+™ Dimethyl Histone H3 (Lys36) set includes a Dimethyl Histone H3 (Lys36) antibody, a Normal Rabbit Serum, and positive control primers which amplify a 87 bp region of the human GAPDH coding region. The Dimethyl Histone H3 (Lys36) antibody and negative controls are supplied in a scalable "per ChIP" reaction size and can be used to functionally validate the precipitation of dimethyl Histone H3 (Lys36) associated chromatin. |
| Alternate Names |
|
| Background Information | Histones are highly conserved proteins that serve as the structural scaffold for the organization of nuclear DNA into chromatin. The four core histones, H2A, H2B, H3, and H4, assemble into an octamer (2 molecules of each). Subsequently, 146 base pairs of DNA are wrapped around the octamer, forming a nucleosome, the basic subunit of chromatin. Histones are modified post-translationally by the actions of enzymes in both the nucleus and cytoplasm. These modifications regulate DNA transcription, repair, recombination, and replication. The most commonly studied modifications are acetylation, phosphorylation, methylation, and ubiquitination. These modifications can alter local chromatin architecture, or recruit trans-acting factors that recognize specific histone modifications (the "histone code" hypothesis). The modifications occur predominantly on the N-terminal and C-terminal tails that extend beyond the nucleosome core particle. Methylation of histone H3 on Lys36 (H3K36me2/3) is tightly associated with actively transcribed genes, and this modification is found primarily within the coding region, suggesting H3K36 methylation is necessary for efficient RNA polymerase II elongation and processivity. |
| Product Information | |
|---|---|
| Format | Serum |
| Control |
|
| Presentation | Anti-Dimethyl Histone H3 (Lys36) (Rabbit Polyclonal), Part No. CS207357. One vial containing 25 µL rabbit polyclonal serum containing 0.05% sodium azide, before the addition of 30% glycerol. Store at -20°C. Normal Rabbit Serum, Part No. CS200585. One vial containing 25 µL antiserum containing 0.05% sodium azide. Store at -20°C. ChIP Primers, GAPDH coding D2. Part No. CS207323. One vial containing 75 μL of 5 μM of each primer specific for human GAPDH coding region. (chr12:6647453+6647539, hg 19 build) Store at -20°C. FOR: 5’ GCC ATG TAG ACC CCT TGA AGA GC 3’ REV: 5’ ACT GGT TGA GCA CAG GGT ACT TTA T 3’ |
| Quality Level | MQ100 |
| Applications | |
|---|---|
| Application | This ChIPAb+ Validated Antibody & Primer Set conveniently includes the antibody, matched IgG negative control antibody & set control PCR primers that detect a known positive locus. |
| Key Applications |
|
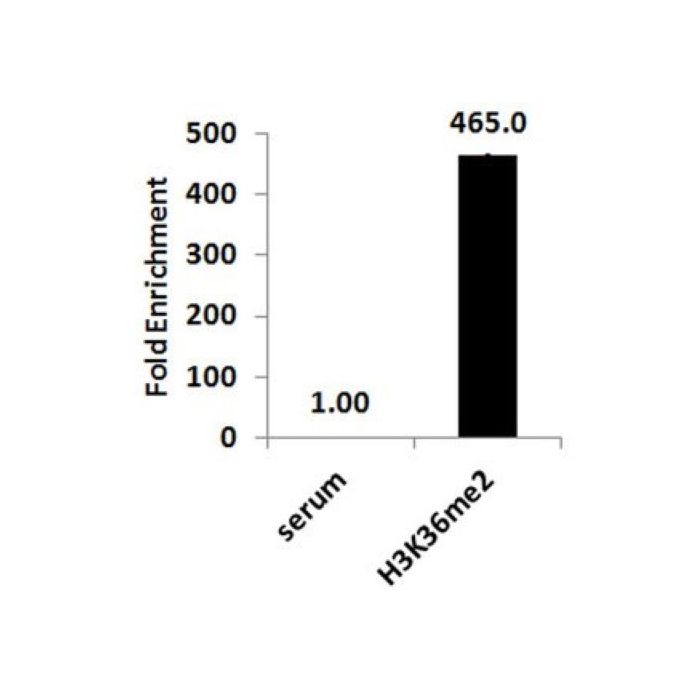
| Application Notes | Chromatin Immunoprecipitation: Sonicated chromatin prepared from HeLa cells (1e5 cell equivalents per IP) were subjected to chromatin immunoprecipitation using 1 µL of either Normal Rabbit Serum (Part No. CS200585), or 1 µL Anti-Dimethyl Histone H3 (Lys36) (Part No.CS207357) and the Magna ChIP™ HiSens (Cat. # 17-10460). Successful immunoprecipitation of dimethyl Histone H3 (Lys36) associated DNA fragments was verified by qPCR using ChIP Primers, GAPDH coding D2 (Part No. CS207323) as a positive locus, and GAPDH promoter (Part No. 22-004) as a negative locus. (Figure 2). Data are presented as percent input of each IP sample relative to input chromatin for each amplicon and ChIP sample as indicated. Please refer to the Magna ChIP™ HiSens (Cat. # 17-10460) or EZ-MagnaChIP™ HiSens(Cat. # 17-10461) protocol for experimental details. Chromatin Immunoprecipitation: Sonicated chromatin prepared from HeLa cells (1e5 cell equivalents per IP) were subjected to chromatin immunoprecipitation using 1 µL of either Normal Rabbit Serum (Part No. CS200585), or 1 µL Anti-Dimethyl Histone H3 (Lys36) (Part No.CS207357) and the Magna ChIP™ HiSens (Cat. # 17-10460). Successful immunoprecipitation of dimethyl Histone H3 (Lys36) associated DNA fragments was verified by qPCR using ChIP Primers, GAPDH coding D2 (Part No. CS207323) as a positive locus, and GAPDH promoter (Part No. 22-004) as a negative locus. Please refer to the Magna ChIP™ HiSens (Cat. # 17-10460) or EZ-MagnaChIP™ HiSens(Cat. # 17-10461) protocol for experimental details. Western Blotting Analysis: A 1:10,000 dilution of this antibody detected dimethyl Histone H3 (Lys36) in 10 µg HeLa acid extract and demonstrated a loss of signal in 0.5 µg of unmethylated recombinant Histone H3 Dot Blot Analysis: Representative lot data. Absurance™ Histone H3 Antibody Specificity Array (Cat. No. 16-667, Fig. A) Absurance™ Histone H2A, H2B, and H4 Antibody Specificity Array (Cat. No. 16-665, Fig. B) were probed with Anti-Dimethyl Histone H3 (Lys36) (1:10,000 dilution). Proteins were visualized using a Donkey Anti-Rabbit IgG secondary antibody conjugated to HRP and a chemiluminescence detection system. |
| Biological Information | |
|---|---|
| Immunogen | KLH-conjugated linear peptide corresponding to a region near the N-terminus of human Histone H3 dimethylated at Lys36. |
| Epitope | Dimethylated Lys36 |
| Host | Rabbit |
| Specificity | Recognizes Dimethyl Histone H3 (Lys36). No detectable cross-reactivity unmethylated, mono, or dimethylated Lys36. No detectable cross reactivity to other histone modifications as measured by AbSurance Histone Antibody Specificity Arrays. |
| Species Reactivity |
|
| Species Reactivity Note | Wide reactivity with other species is expected. |
| Antibody Type | Polyclonal Antibody |
| Entrez Gene Number |
|
| Entrez Gene Summary | Histones are basic nuclear proteins that are responsiblefor the nucleosome structure of the chromosomal fiber in eukaryotes. Nucleosomes consist of approximately 146 bp of DNA wrapped around a histone octamer composed of pairs of each of the four core histones (H2A, H2B, H3, and H4). The chromatin fiber is further compacted through the interaction of a linker histone, H1,with the DNA between the nucleosomes to form higher order chromatin structures. This gene is intronless and encodes a member of the histone H3 family. Transcripts from this gene lack polyA tails; instead, they contain a palindromic termination element. This gene is located separately from the other H3 genes that are in the histone gene cluster on chromosome 6p22-p21.3. [provided by RefSeq] |
| Gene Symbol |
|
| Modifications |
|
| UniProt Number |
|
| UniProt Summary | FUNCTION: Core component of nucleosome. Nucleosomes wrap and compact DNA into chromatin, limiting DNA accessibility to the cellular machineries which require DNA as a template. Histones thereby play a central role in transcription regulation, DNA repair, DNA replication and chromosomal stability. DNA accessibility is regulated via a complex set of post-translational modifications of histones, also called histone code, and nucleosome remodeling. SUBUNIT STRUCTURE: The nucleosome is a histone octamer containing two molecules each of H2A, H2B, H3 and H4 assembled in one H3-H4 heterotetramer and two H2A-H2B heterodimers. The octamer wraps approximately 147 bp of DNA. SUBCELLULAR LOCATION: Nucleus. TISSUE SPECIFICITY: Expressed in testicular cells. Developmental stage Expressed during S phase, then expression strongly decreases as cell division slows down during the process of differentiation. PTM: Acetylation is generally linked to gene activation. Acetylation on Lys-10 impairs methylation at Arg-9. Acetylation on Lys-19 and Lys-24 favors methylation at Arg-18 By similarity. Citrullination at Arg-9 and/or Arg-18 by PADI4 impairs methylation and represses transcription By similarity. Asymmetric dimethylation at Arg-18 by CARM1 is linked to gene activation. Symmetric dimethylation at Arg-9 by PRMT5 is linked to gene repression By similarity. Methylation at Lys-5, Lys-37 and Lys-80 are linked to gene activation. Methylation at Lys-5 facilitates subsequent acetylation of H3 and H4. Methylation at Lys-80 is associated with DNA double-strand break (DSB) responses and is a specific target for TP53BP1. Methylation at Lys-10 and Lys-28 are linked to gene repression. Methylation at Lys-10 is a specific target for HP1 proteins (CBX1, CBX3 and CBX5) and prevents subsequent phosphorylation at Ser-11 and acetylation of H3 and H4. Methylation at Lys-5 and Lys-80 require preliminary monoubiquitination of H2B at Lys-120. Methylation at Lys-10 and Lys-28 are enriched in inactive X chromosome chromatin By similarity. Phosphorylated at Thr-4 by GSG2/haspin during prophase and dephosphorylated during anaphase. At centromeres, specifically phosphorylated at Thr-12 from prophase to early anaphase. Phosphorylated at Ser-11 during the whole mitosis. Phosphorylation at Ser-11, which is linked to gene activation, prevents methylation at Lys-10 but facilitates acetylation of H3 and H4. Phosphorylated at Ser-29 by MLTK isoform 1, RPS6KA5 or AURKB during mitosis or upon ultraviolet B irradiation By similarity. Phosphorylation at Ser-11 is crucial for chromosome condensation and cell-cycle progression during mitosis and meiosis. In addition phosphorylation at Ser-11 is important during interphase because it enables the transcription of genes following external stimulation, like stress or growth factors. Phosphorylation at Ser-11 is also an essential regulatory mechanism for neoplastic cell transformation. Phosphorylation at Ser-11 by AURKB/Aurora-B mediates the dissociation of HP1 proteins (CBX1, CBX3 and CBX5) from heterochromatin. Ubiquitinated By similarity. SIMILARITY:Belongs to the histone H3 family. |
| Molecular Weight | ~17 kDa |
| Product Usage Statements | |
|---|---|
| Quality Assurance | Sonicated chromatin prepared from HeLa cells (1e5 cell equivalents per IP) were subjected to chromatin immunoprecipitation using 1 µL of either Normal Rabbit Serum (Part No. CS200585), or 1 µL Anti-Dimethyl Histone H3 (Lys36) (Part No.CS207357) and the Magna ChIP™ HiSens Kit (Cat. # 17-10460). Successful immunoprecipitation of dimethyl-Histone H3 (Lys36) associated DNA fragments was verified by qPCR using ChIP Primers, GAPDH coding D2 (Part No. CS207323). Please refer to the Magna ChIP™ HiSens (Cat. # 17-10460) or EZ-Magna ChIP™ HiSens (Cat. # 17-10461) protocol for experimental details. |
| Usage Statement |
|
| Storage and Shipping Information | |
|---|---|
| Storage Conditions | Stable for 1 year at -20°C from date of receipt. Handling Recommendations: Upon first thaw, and prior to removing the cap, centrifuge the vial and gently mix the solution. Aliquot into microcentrifuge tubes and store at -20°C. Avoid repeated freeze/thaw cycles, which may damage IgG and affect product performance. |
| Packaging Information | |
|---|---|
| Material Size | 25 assays |
| Material Package | Recommended use: 1 μL of antibody per chromatin immunoprecipitation (dependent upon biological context). |